Recent Studies
A novel chemical strategy development at Dr. Muhammad Jbara research group will allow deep understanding of the gene expression processes
Deciphering the Role of the Ser-Phosphorylation Pattern on the DNA-Binding Activity of Max Transcription Factor Using Chemical Protein Synthesis
Raj V. Nithun, Yumi Minyi Yao, Xiaoxi Lin, Shaimaa Habiballah, Ariel Afek, and Muhammad Jbara
Transcription factors are important proteins that control the rate of transcription of genetic information. These proteins regulate gene expression by selectively interacting with certain DNA sequences to activate or suppress specific genes by "on and off" mechanisms. A transcription factor called Max plays a central role in regulating 15% of human genes. This protein is involved in several key cellular processes, including cell division, cell differentiation and cell death, and therefore, dysregulation to its activity is involved in many diseases, including cancer (70%). Understanding the function of transcription factors such as Max and their binding to DNA is a critical step for understanding gene expression processes and how failure in this process contributes to the development of diseases. Many studies have suggested that structural changes on the surface of transcription factors affect their operation and normal function in the cell. For example, many proteins are activated or deactivated by the addition or removal of a phosphate group in a process called phosphorylation. The main barrier to gaining molecular insights into the effect of phosphorylation on transcription factors such as Max is the difficulty in obtaining homogeneous Max samples with phosphorylation desired patterns using conventional biological methods.
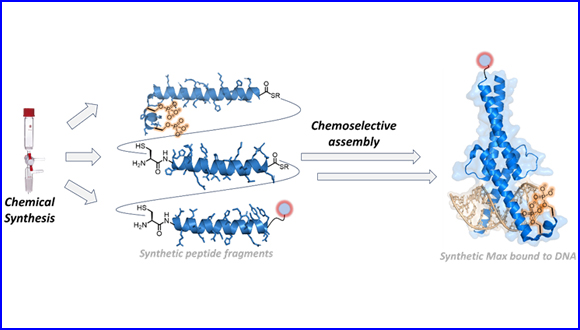
The breakthrough in this research is based on the researchers' success in preparing synthetic proteins that are identical to the natural transcription factors in the cell and thus studying their activity in a controlled and precise manner. In this study, the authors developed a novel chemical strategy to construct a library of transcription factors to investigate the effect of natural modifications such as phosphorylation on the activity of Max. The researchers combined solid phase synthesis and chemoselective ligation chemistry to build synthetic Max analogues, while introducing modifications such as phosphorylation and fluorescent markers and proved by biochemical analysis that the synthetic proteins they prepared are identical to the natural proteins in the cells. This development made it possible to understand the activity of the Max protein, and specifically the effect of phosphorylation on the recognition and binding with essential DNA sites. The authors found that phosphorylation affects the binding efficiency of Max to DNA, and it even disrupted the interaction to DNA. In addition, they discovered that the position of the phosphorylation site is critical for the activity of Max and its binding to DNA. Also, by scanning the binding of the protein to thousands of DNA sequences, by using high throughput screening technology, they showed that despite disrupting the connection to the DNA, the phosphorylation does not change Max's preference for the specific DNA sequences.
The development of synthetic techniques to create proteins such as transcription factors is essential for the characterization and understanding of complex biological mechanisms. We anticipate that synthetic developments from this study will pave the way for other transcription systems. In addition, this research is a basis for the understanding of additional modifications beyond phosphorylation. Beyond that, we anticipate that the methods developed in this work will be important for the engineering and development of innovative molecules that can be used to increase and suppress gene expression and thereby contribute to other fields of research in biology and medicine.